Thank you! A giant leap forward. Looking good. Your updated information and tools have already supplied me with 2 new relatives second cousins once removed, and your chromosome browser allowed me to determine which segments on which chromosomes we have in common. Looking forward to when you can implement a “1-to-many” type of interface where you can compare more than one match in a chromosome browser. Here’s hoping:)

We’re excited to announce major updates and improvements to DNA Matching rolled out today for all our users. Anyone who took a MyHeritage DNA test, and anyone who uploaded DNA data from another service, will now receive more accurate DNA Matches; more plentiful matches (about 10x more); fewer false positives; more specific and more accurate relationship estimates; and indications on lower confidence DNA Matches to help focus research efforts. We also added the long-requested Chromosome Browser feature, described below.
These improvements have been many months in the making by our Science Team. They took much time and effort because we wanted to perfect the science and provide our users with optimal results.
What is DNA Matching?
MyHeritage DNA currently has more than one million people in the DNA database. 1.075 million to be precise. DNA Matching compares DNA kits in the MyHeritage database to each other, to find relatives, i.e. individuals who share DNA segments with each other, and to help explain how these individuals are related. The presence of shared DNA segments between two people can indicate a blood relationship, meaning that the shared segments were inherited from a common ancestor. If the shared segments are numerous and large, a blood relationship is more certain. On the other hand, if the shared segments are small in number and size, it can also be a matter of coincidence, indicating no blood relationship at all. When a match is reported that is not a relative at all, it is a false positive.
If you have taken a MyHeritage DNA test and received the results, or uploaded your DNA data to MyHeritage, then you will have received a list of your DNA Matches. The matches are updated daily, and users are notified by a weekly email about the best new matches they received that week. By “best” we mean the matches having the largest amount of shared DNA indicating a closer relationship. The list of DNA Matches shows individuals who share DNA segments with you, the amount and percentage of DNA you share, the number of DNA segments you share, and the size of the largest shared segment. MyHeritage also estimates the relationship by analyzing the number and size of the shared DNA segments in each match and comparing them to a reference pool of hundreds of thousands of other matches with known relationships according to family trees that were confirmed by DNA. The DNA Match Review Page offers leads that you can follow up on to trace your lineage back to your common ancestor.
As of today, users who have received DNA Matches before, will see modified and enhanced matches following these improvements. This means many new matches will appear. Some matches that existed before, that were false positives, will disappear. Many matches will have their parameters changed (e.g. amount of shared DNA) to more accurate values. Users who have not received matches yet, will receive the higher quality matches from day one.
How does DNA Matching work?

Let’s start with a quick overview of how DNA Matching works. Then we’ll delve into the improvements we’ve made in the different stages of the process.
The process begins when you take a DNA test and send your sample to our lab. In the lab, we read your DNA and produce a data file with the information. We don’t read every part of your DNA, which amounts to about 3 billion points. This is an expensive method called whole genome sequencing, currently reserved for specific clinical and research applications. Instead we focus on reading approximately 700,000 locations in your DNA that are known to vary between individuals, called single nucleotide polymorphisms (SNPs, pronounced “snips”). This method is called genotyping and it produces a data file that lists each SNP that we read, its position in your DNA, and the two genotypes we found there (i.e. the A, T, G, or C you inherited from each parent). If you uploaded DNA data from another service, we receive the data file with the same information.
Next, comes phasing. In every pair of chromosomes, every person gets one chromosome from their mother and one from their father. The genotyping technology that reads your DNA sample determines what genotypes you inherited from your parents for each SNP, but it doesn’t tell us which groups of variants were co-inherited from the same parent. Phasing helps us sort this out. It clusters all variants inherited from one of your parents in one bucket and the variants inherited from the other parent in another bucket.
After phasing we use imputation to infer the SNPs we did not read. Think of imputing DNA as reading a sentence with some of the letters missing — there’s a good chance that you can infer the missing letters from context. Not all DNA service providers read the same SNPs. To find DNA Matches for individuals who used different DNA service providers, it is important to infer the SNPs that were not read before comparing results. Some people question the accuracy of imputation. However, we found that this method is very accurate when used properly, and in some situations its usage is inevitable.
The next step is to perform the actual matching, and compare all the DNA kits in the database that have not been opted out of matching by their owners, to each other. We do this on a very scalable system called Hadoop that allows us to perform massive distributed processing very efficiently. Matching identifies the shared segments between each pair of kits, from which the relationship of the two individuals (if any) can be deduced. Adjacent shared segments are then “stitched” if they are considered contiguous.
Finally, we use advanced statistical algorithms called classifiers to review the DNA Matches and reject false positives, to determine the confidence level of the matches that were not rejected, and to suggest the type of relationship for each match. That’s how we create your list of DNA Matches.
How have we improved DNA Matching?
We fixed the phasing. The previous processing of DNA Matches had occasional errors in the phasing stage. These errors caused some false positives where we previously over-estimated shared segments of very distant relatives. It also caused problems where we previously under-estimated shared segments of close relatives. We now use a better algorithm that fixes these phasing errors.
We have improved the accuracy of our imputation significantly by increasing the number of reference genomes more than tenfold. Just like reading 10 times as many books would enable a person to accurately infer missing letters from more sentences, increasing our reference genome panel greatly increased our ability to impute SNPs we did not read, more accurately.
In the matching stage we have re-calibrated the threshold for genotyping errors. The technology that reads your DNA sample makes occasional mistakes. These are called genotyping errors. If a genotyping error occurs in the middle of what should be a shared segment between DNA Matches, that segment will not appear to be identical, and it may be split into two matching segments that are smaller. We re-calibrated the threshold for when we ignore small mismatches between otherwise matching segments, and instead treat the shared segments as identical despite small pieces that do not match. This method compensates for unavoidable genotyping errors. If we ignore mismatched sections that are too large, we will accidentally assume a segment is shared when it really isn’t; if we don’t ignore mismatched sections that are the result of genotyping errors, we are likely to miss real DNA Matches. The new calibration is stricter than the previous one, which means that fewer false positives will slip through.
More distant matches are now allowed. After increasing the accuracy of the matches and calibrating the above parameters, we felt comfortable allowing more distant matches to be presented to you. Previously the minimum of shared DNA for a match was 12 cM and now the minimum is 8 cM. This together with the other improvements yielded a tenfold increase in the number of DNA Matches our users will now receive.
These matches will appear automatically for anyone who already took a MyHeritage DNA test or anyone who has already uploaded their DNA to MyHeritage in the past and for anyone who does so in the future.
Better stitching of adjacent segments. In addition to compensating for genotyping errors within segments, it’s necessary to compensate for the remaining phasing errors between segments. For example, excluding the gender chromosome, a mother and daughter are expected to have 22 matching segments in an autosomal DNA test: One entire chromosome from each of the daughter’s chromosome pairs was inherited from her mother, and so each of the 22 autosomal chromosomes should appear to be a single, long, matching segment. However, due to phasing errors, sometimes small sections of the chromosome inherited from the mother are computationally swapped with the parallel sections inherited from the father. This is a result of technical errors, not biological processes. We overcame these errors by increasing the size of the gaps we stitch, while calibrating this precisely to avoid introducing new errors.
The last step of DNA Matching is filtering out false positives and estimating the specific relationship between two individuals with shared DNA segments. Because many of us are descendants of the same very ancient ancestors, we often have tiny shared DNA segments with individuals we wouldn’t really consider family. We sought a method to filter out such matches that only frustrate genealogists. To this end, we measure false positives internally by looking at trios — these are sets of child, mother, and father who were all tested with MyHeritage DNA kits, and received results that validated that the relationships between the parents and child is correct. Any match that a child has with another individual, who does not match neither the father nor the mother is suspected to be a false positive and is called a child-only match. We measure the percentage of child-only matches among all matches that are returned for children in all known trios on MyHeritage, and this figure is called the percentage of suspected false positives indicated by child-only matches. We managed to bring this figure down to 16–20 percent, which is a good result that as far as we know is equivalent to or better than all other DNA services. Our improved classifier algorithms have succeeded in bringing our false positive rate to an all-time low.
But we did not stop there. We wanted to create a method that lets you focus your genealogy searches in the most effective way. For this, we used our statistical algorithms to categorize the matches into: high, medium, and low confidence matches. Matches that are low or medium confidence are labeled as such in the website. These are DNA Matches that should be treated with skepticism because they are at risk of being false positives. Such matches typically have very few, very small shared DNA segments. These indications allow you to make the best use of your time. Follow up first on high confidence matches, and if you are in the mood for a challenge, sift through lower and medium confidence matches for hidden treasures. Note that low and medium confidence matches are excluded from the weekly notification emails on new matches.
The new classifiers are so good, that the percentage of child-only matches that are not flagged as low or medium confidence, is now less than 5%. In other words, whenever you review a DNA Match on MyHeritage that is not marked as low confidence or medium confidence, you can now be almost certain that you are not wasting your time on a false positive. If the match you are reviewing is estimated to be a second cousin or closer, there is so much shared DNA that you can rest assured it is not a false positive.
The accuracy of estimating a relationship of a DNA Match is measured using two parameters called recall and precision. Perfect accuracy means both telling a user the correct relationship to a DNA Match every single time (recall), while suggesting only that relationship and not proposing a wider range of possible relationships (precision). For example, if two individuals are in fact siblings, a perfect algorithm will suggest that they are siblings and only suggest that they are siblings; it will not estimate that they are either siblings or cousins. (However, biologically, due to the nature of DNA inheritance, this theoretical perfect algorithm isn’t possible). MyHeritage can now suggest DNA Matches’ correct relationship 93% of the time for distant relatives like 4th and 5th cousins which is incredibly difficult to do. For closer relatives, the accuracy is much higher and is close to 100%. At the same time, we will only suggest about 2 or 3 possible relationships for DNA Matches who are 1st cousins or closer. For distant cousins we will show an average of up to 5 possible relationships (such as second cousin once removed and third cousin) — a relatively narrow range, considering. The precision and recall of MyHeritage’s relationship estimates are now much better than before.
We validated the high quality of our new DNA Matching algorithm by comparing new DNA Match lists to those produced by other DNA companies and the results are very similar.
Certain particularly endogamous populations, like Ashkenazi Jews, pose a unique DNA Matching challenge. Because these populations experienced significant intermarriage, non-genealogically related individuals within these population have more shared DNA than would be expected for non-relatives otherwise. MyHeritage trained an additional classifier algorithm using machine learning to classify Ashkenazi relationships with higher resolution than any other DNA service. We used this classifier to provide better rejection of false positives for Ashkenazi Jews, bringing them to the same level of false positives as the general population.
What do these improvements mean for MyHeritage DNA users?
Now, you will get:
- More accurate DNA Matches
- About 10 times as many DNA Matches
- More specific and more accurate relationship estimates
- Indications of DNA Match confidence levels to help focus your research efforts
Chromosome Browser
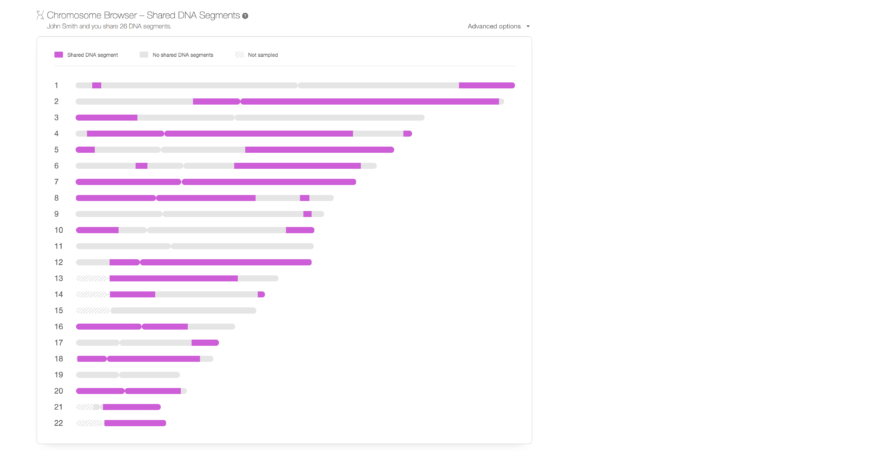
Along with the improvements in accuracy, we’ve also added new features to enhance the use of DNA Matches. The first, by popular demand, is a Chromosome Browser for shared DNA Matches. It was added to the DNA Match Review page.
A chromosome browser is a schematic representation of a person’s chromosomes where DNA segments can be visualized. Many of our users have requested a chromosome browser and we know this is an important tool for genealogists. Hence, we promised that we’d develop one and now we have delivered on this promise. The new Chromosome Browser on MyHeritage is an initial release, that will be enhanced further soon, intended for viewing shared DNA segments for any DNA Match. It’s a free feature that can be used by all users on MyHeritage who have taken the DNA test or uploaded DNA data. It shows the shared segments between you and a DNA Match in purple. When you hover your mouse over any shared segment you can see the genomic position of the shared segment, the size of the segment, and the number of SNPs there. Grey segments are not shared with the DNA Match and crisscrossed sections were not analyzed due to the lack of SNPs in those regions. Note that although we’d never ever allow another user to download your raw DNA data, if another user has a shared segment with you and can view its details (position and size), then by reviewing that segment’s information on his or her own DNA, that other user can deduce the genotypes that you have in your DNA in that particular segment. Users who prefer to prevent other users who match their DNA from viewing the details of shared segments can opt out of this feature by using a new privacy setting that we added for this purpose.
The Chromosome Browser also includes the ability to download data about shared segments. This is accessible via the “Advanced Options” menu at the top right corner of the Chromosome Browser. Advanced users can use this option to download the information about the shared segments and then use it for viewing in other tools or chromosome browsers. More features will be coming out soon, such as the ability to view three or more DNA Matches’ shared segments in the Chromosome Browser simultaneously. Viewing the shared segments of multiple DNA Matches at the same time helps you trace back and identify the shared ancestor who passed down the segment to all the DNA Matches who share it, and then discover how you are all related. We’re also planning to add the ability to print the shared segments displayed by the Chromosome Browser soon.
With some practice, we’re confident that our users will be able to use the new Chromosome Browser feature to start identifying specific segments in their DNA and the ancestor they originated from, gain better insights about their DNA Matches, and better understand the relationship path connecting these DNA Matches. We hope this will help our community break through brick walls, trace their ancestry, and understand how they are related to family members discovered via the DNA Matches.
Facelift and easier navigation
As part of this update we made small revisions to the user interface of the DNA Matches for better consistency with the other DNA screens. Most of these changes are small and barely noticeable, such as buttons on the list of DNA Matches being purple instead of orange. However, one more meaningful improvement is that the DNA Match list page now docks the details of the DNA kit you are viewing at the top, so when you scroll down through the list you will never lose track of whose matches you’re looking at.
Work in progress
Our work is not complete. DNA Matching is always work in progress and will be constantly improved by us in the future. The growing size of our DNA database, as well as the increased association between DNA kits and family trees, provide us with more opportunities to optimize the DNA Matching algorithms, and we intend to do this regularly and improve accuracy further.
With regards to DNA data uploaded from other services, we still do not support DNA Matching to DNA kits that are based on the Illumina GSA chip. These include kits from 23andMe (recent V5 version) and Living DNA. We have support for DNA data from GSA chips working in our lab; it is in fairly good shape, but still not perfect, so we decided to exclude it from this release until it is perfected. This will be added in the next few months.
Ethnicity estimates are separate from DNA Matching and the improvements described here do not affect ethnicity estimates. We are planning an update to our ethnicity reports in the next few months to improve their accuracy too. Stay tuned!
Next steps
Order a MyHeritage DNA kit to benefit from these new features and improvements, if you haven’t done so already. If you have already tested yourself, consider getting DNA kits for some of your relatives, especially older ones, to find more relatives and triangulate your own matches. For example, by testing a cousin of your mother or father, any matches with new relatives you already have that are also shared with that cousin, can be triangulated to a common ancestor, through a path to a more recent ancestor in your family tree that you share with that cousin. The new Chromosome Browser will come in handy for understanding those matches. So if there is a particular branch in your family tree that you are most interested in exploring further, buy additional DNA kits for older relatives of yours in that branch.
If you have already tested your DNA elsewhere, upload your DNA data to MyHeritage. MyHeritage is the only one of the three largest DNA services to support DNA data upload. Take advantage of this while it’s still free and get free DNA Matches and free Ethnicity Estimates for your existing data. With MyHeritage’s substantial DNA database of more than one million people, most of whom tested only on MyHeritage, doing this is a no-brainer. You’ll get your results for free in 1–2 days or less.
If you already manage more than one DNA kit on MyHeritage, please take the time to verify that each kit you manage is associated with the right person. This can be fixed, if need be, by using the “Manage DNA kits” page, accessible from the DNA menu. Use the option “Re-assign kit to a different person” if you have a kit that is not associated properly. This is necessary if all the kits you uploaded for several relatives are still associated with your own family tree profile.
Finally, DNA kits on MyHeritage are much more useful when there is a family tree associated with them. This allows you to get better insights about your DNA Matches, for example by the presence of Smart Matches, shared ancestral surnames, or shared ancestral birth places between you and any DNA Match. If you have a DNA kit on MyHeritage, but no family tree or a very small family tree, now’s a good time to create a family tree or enhance your existing family tree. It will benefit your DNA Matches, but primarily, it will benefit you.
Enjoy,
The MyHeritage Team








Jason Lee
January 11, 2018
Great job! The upgrades are very good.